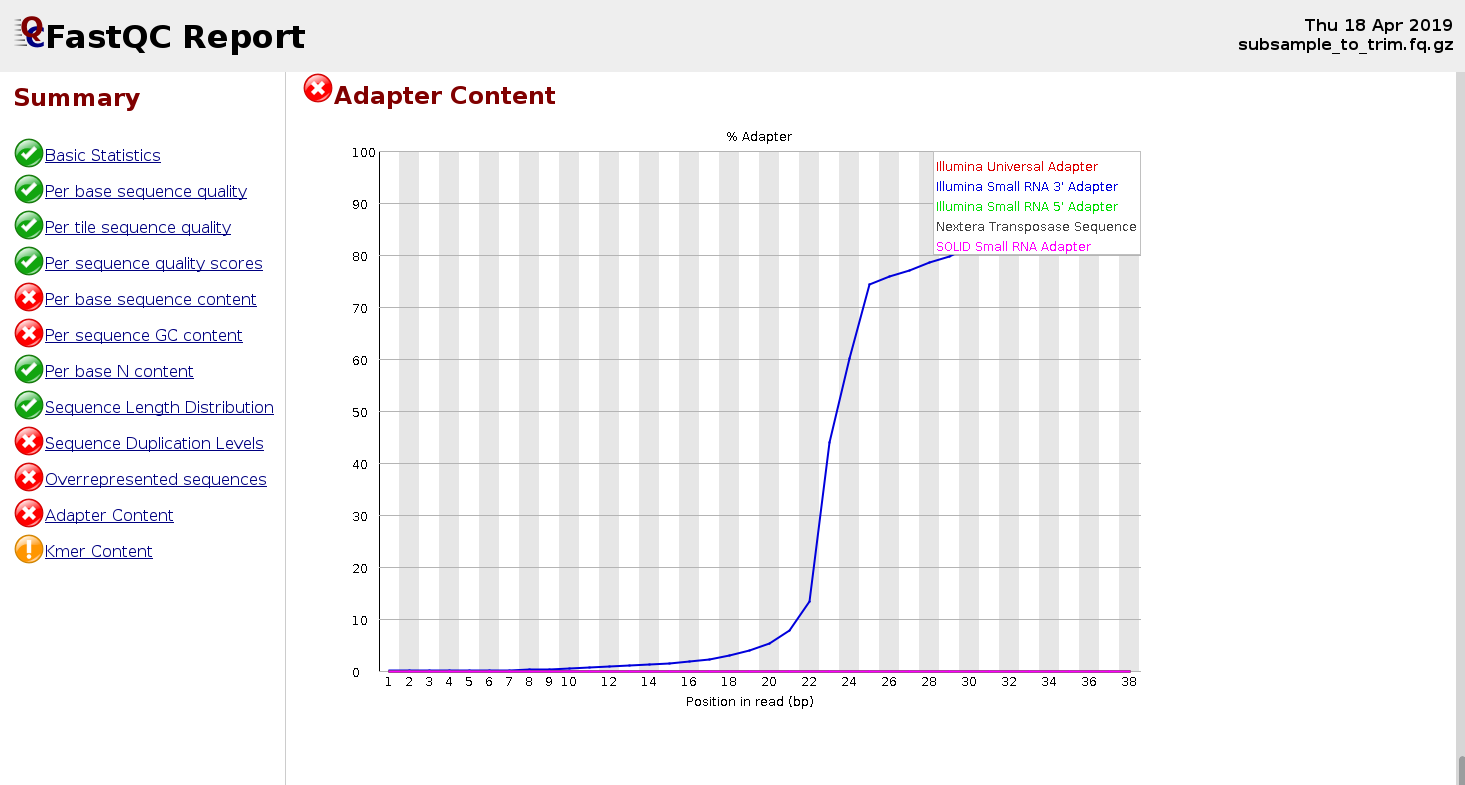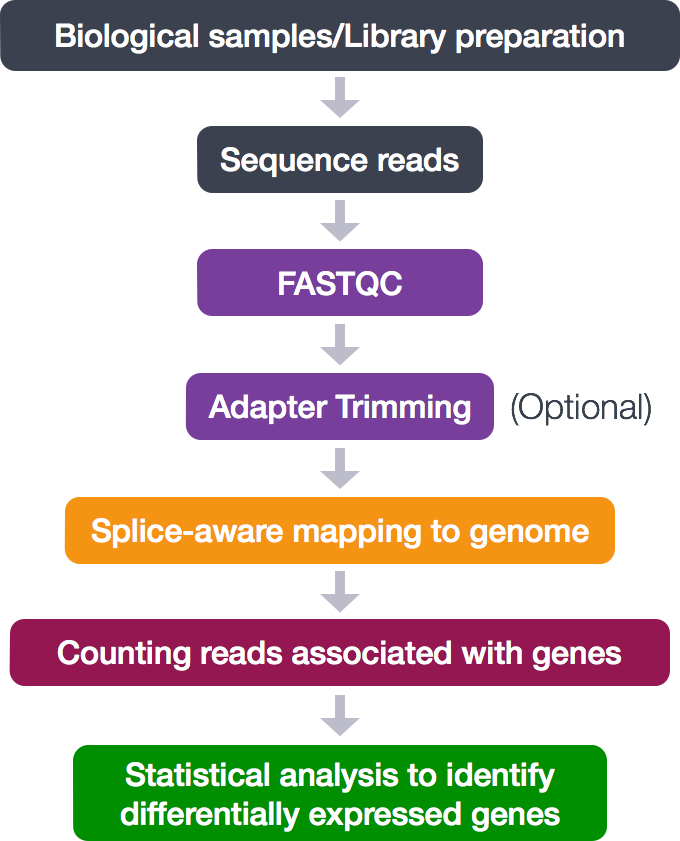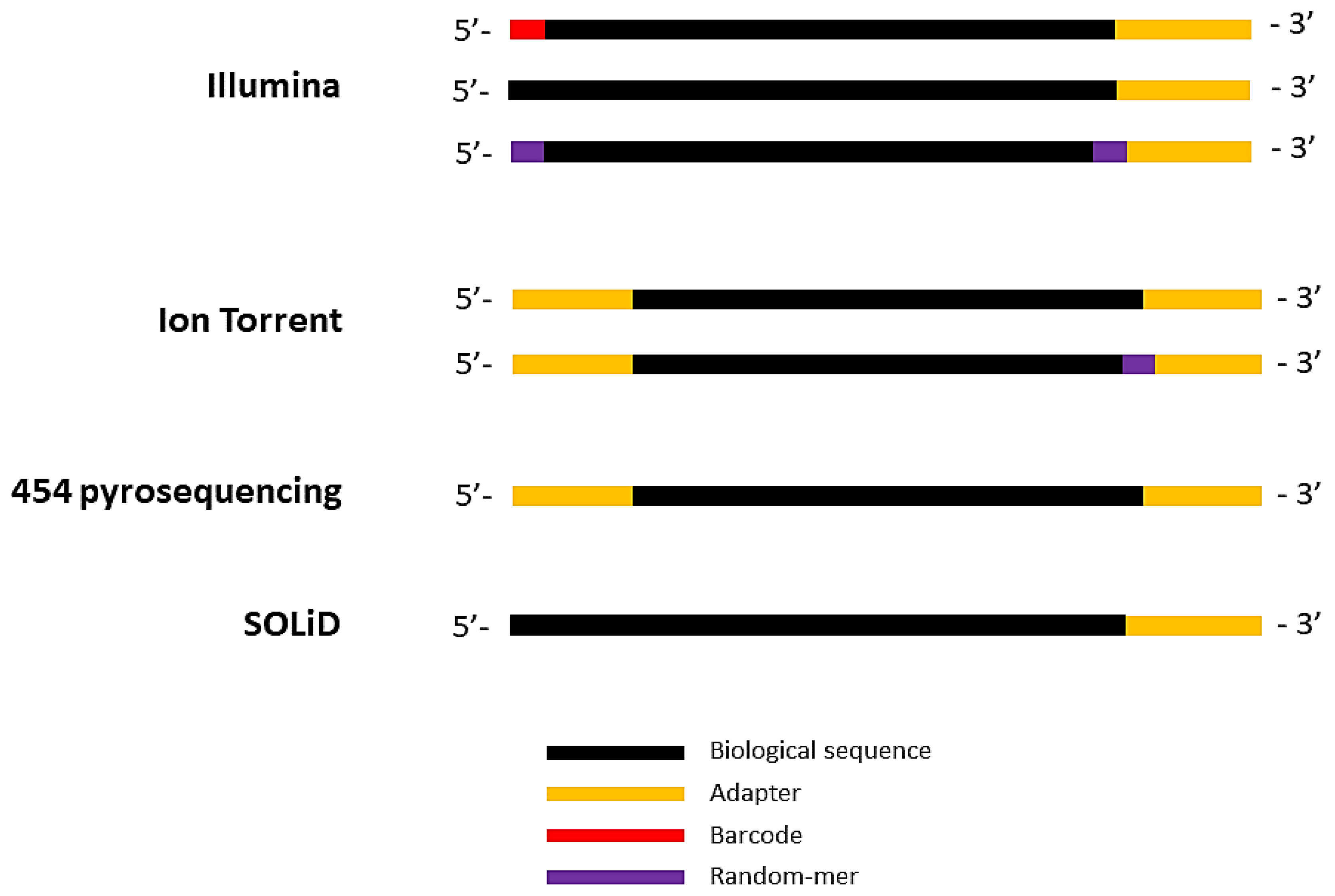
Biomolecules | Free Full-Text | High-Throughput Identification of Adapters in Single-Read Sequencing Data | HTML

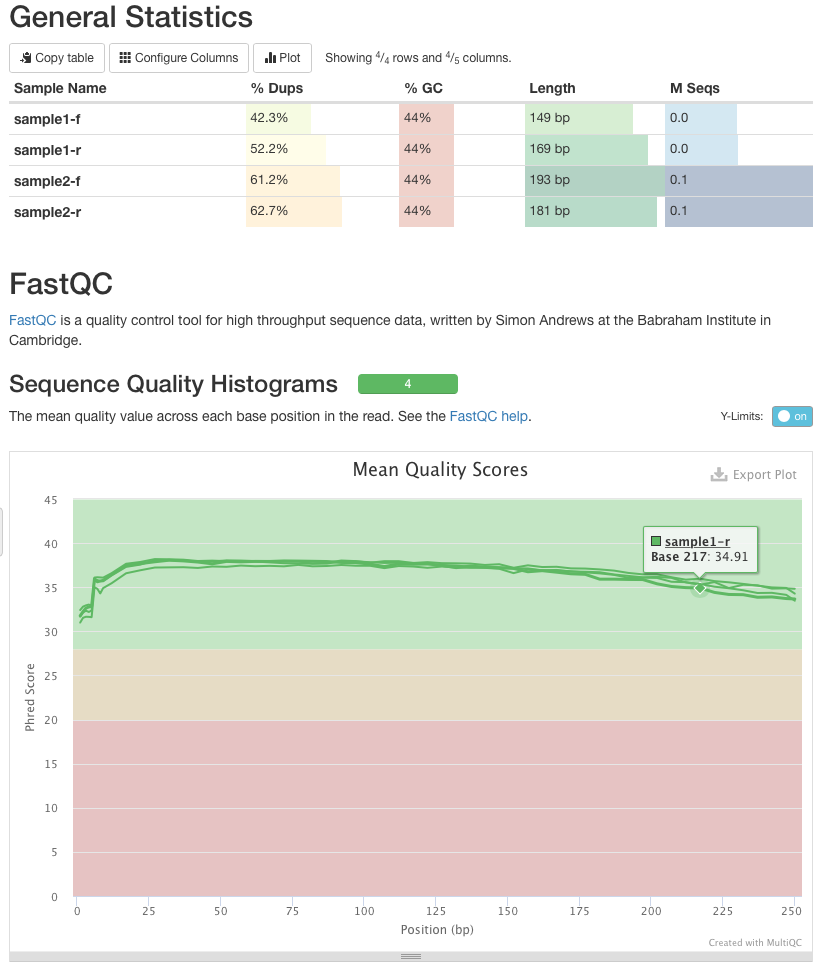
RNA sequencing quality details. FastQC report illustrating the average... | Download Scientific Diagram
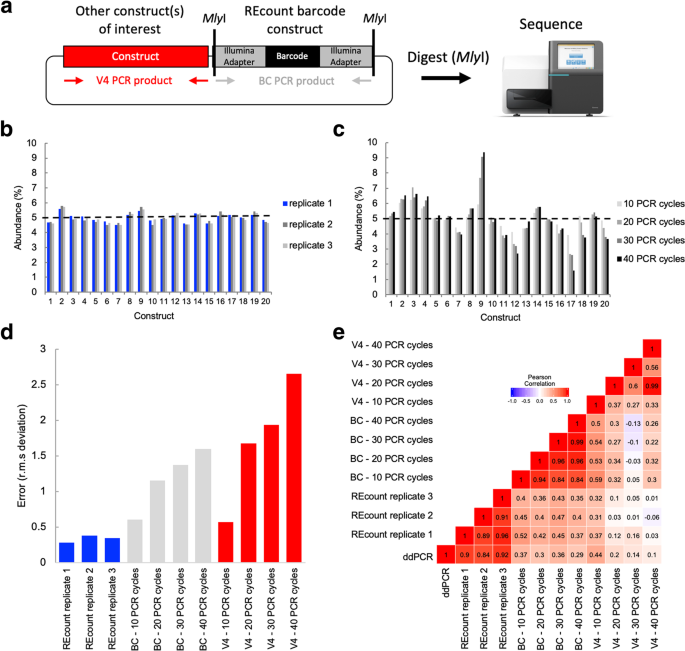
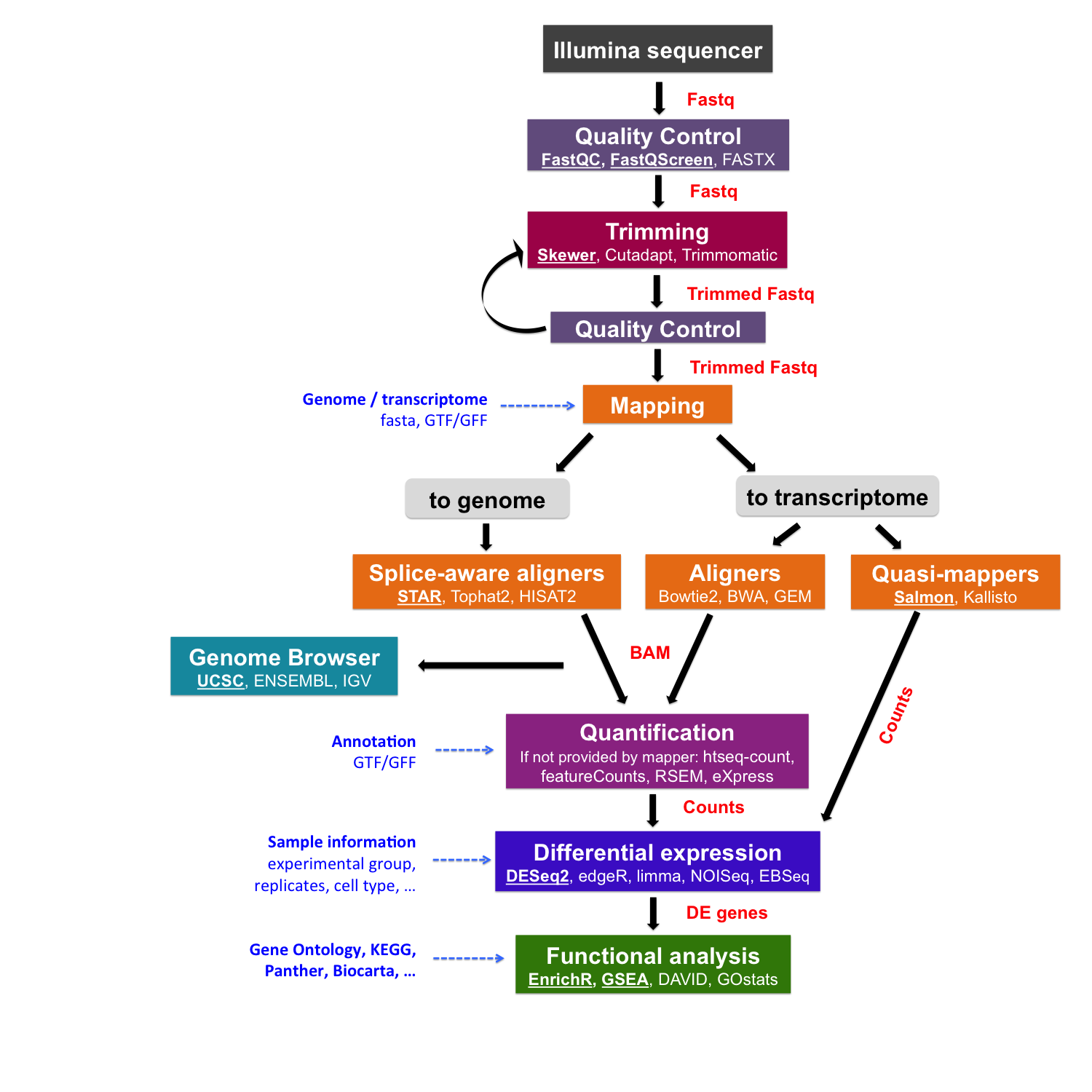
Measuring sequencer size bias using REcount: a novel method for highly accurate Illumina sequencing-based quantification | Genome Biology | Full Text
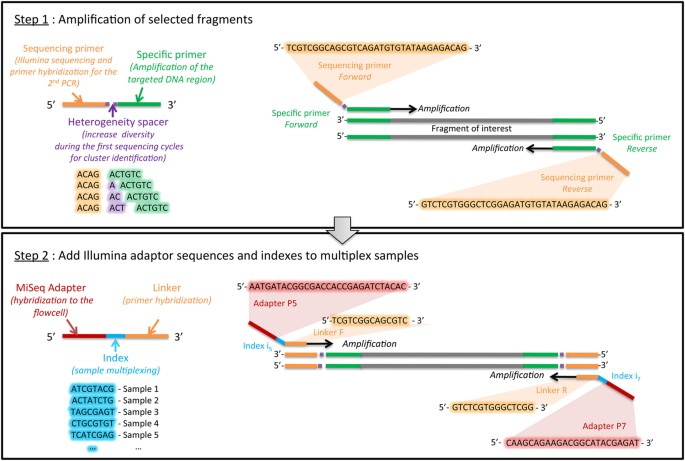
High-throughput sequencing of multiple amplicons for barcoding and integrative taxonomy | Scientific Reports
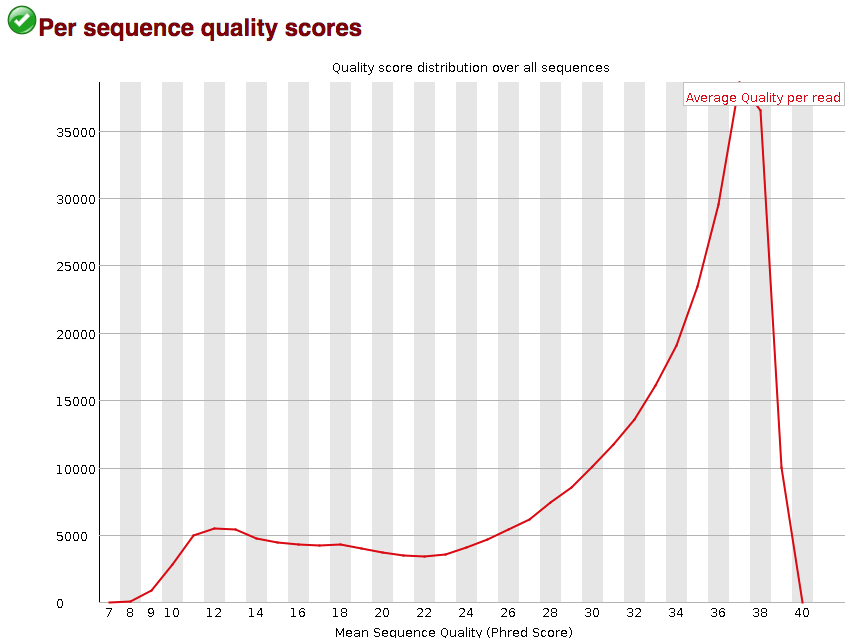
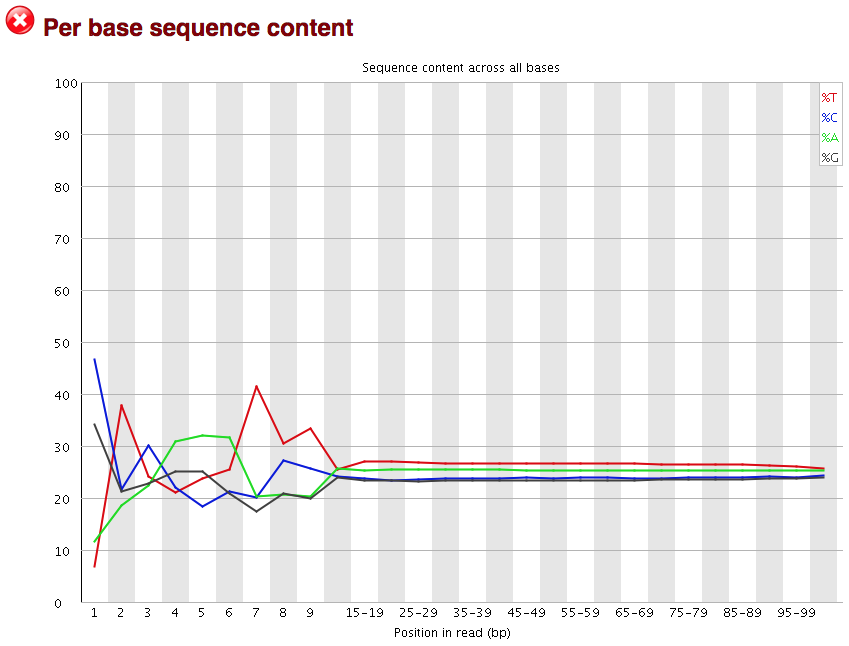
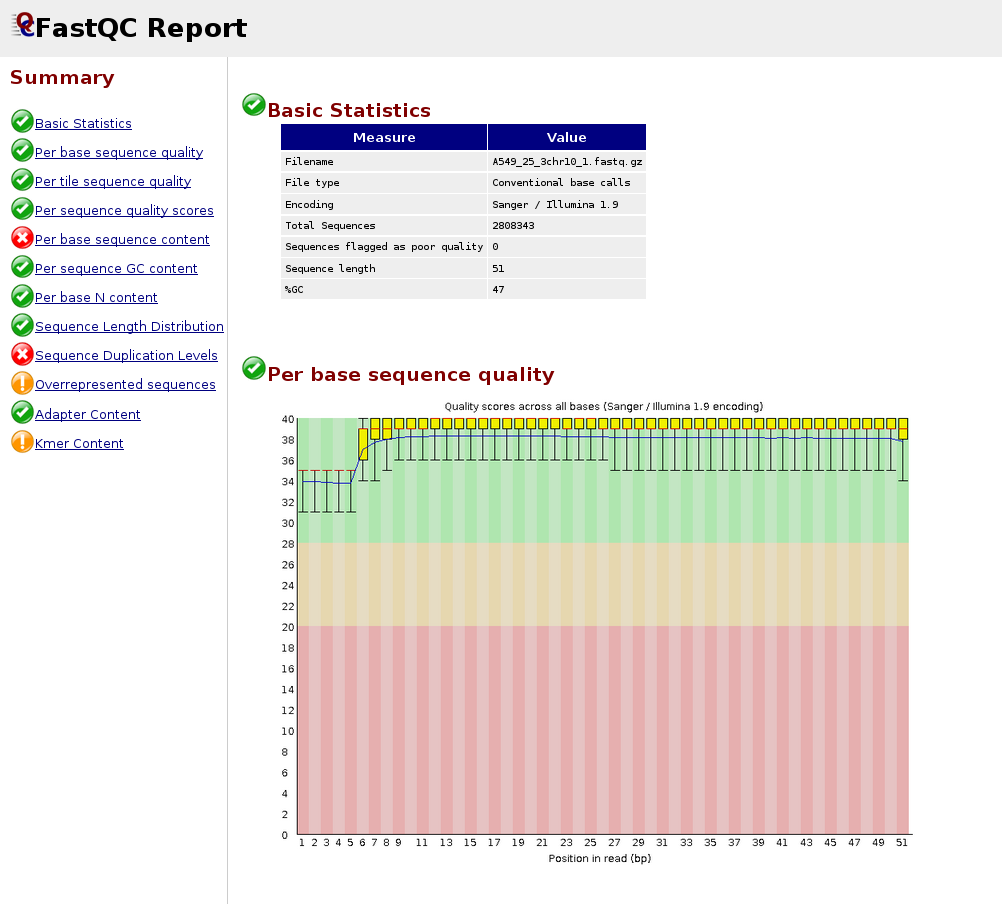
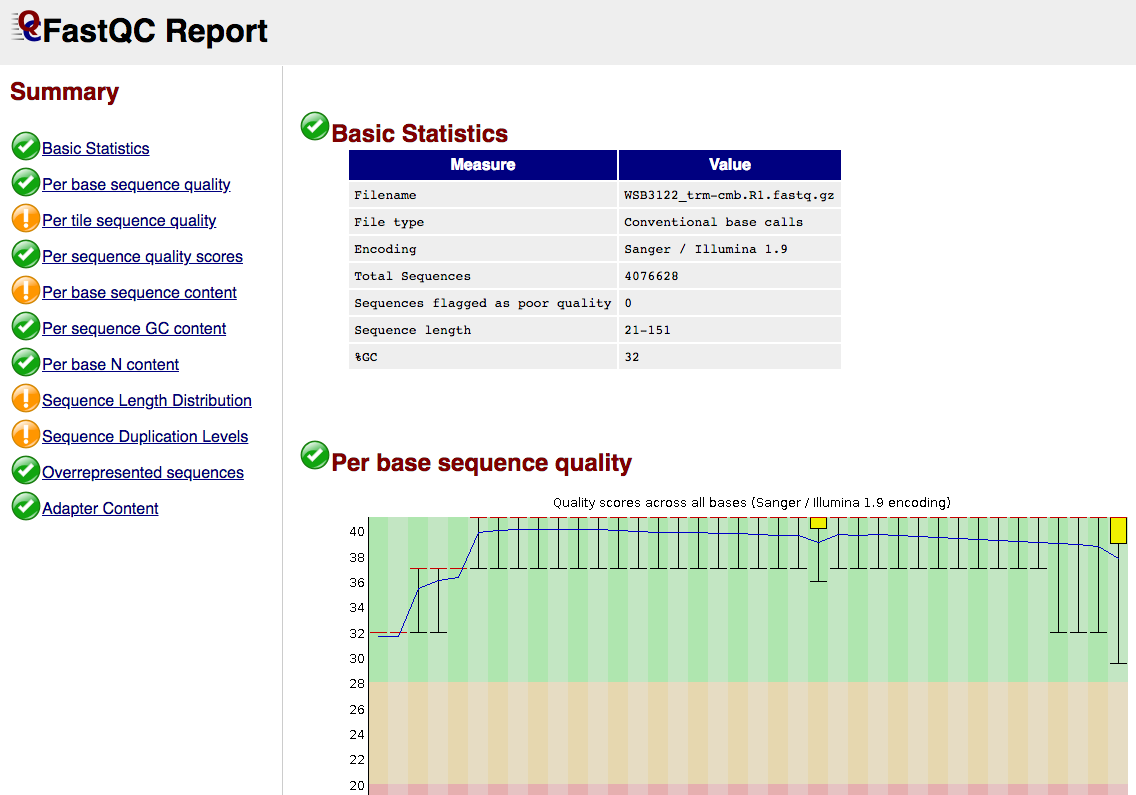
Quality control: Assessing FASTQC results | Introduction to RNA-Seq using high-performance computing - ARCHIVED
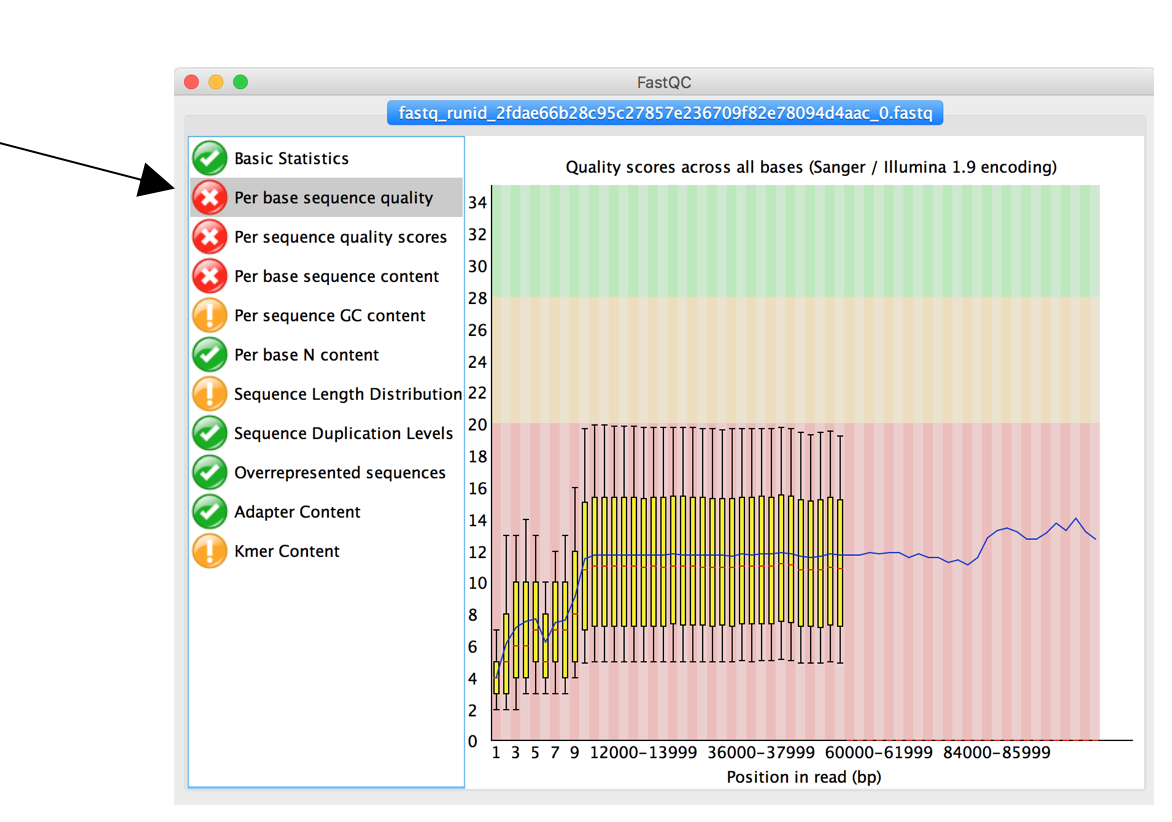
![Introduction to Next Generation Sequencing Bioinformatics”] | [“A Tufts University Research Technology Workshop”] Introduction to Next Generation Sequencing Bioinformatics”] | [“A Tufts University Research Technology Workshop”]](https://rbatorsky.github.io/intro-to-ngs-bioinformatics/img/fastqc_before_trim.png)
Introduction to Next Generation Sequencing Bioinformatics”] | [“A Tufts University Research Technology Workshop”]
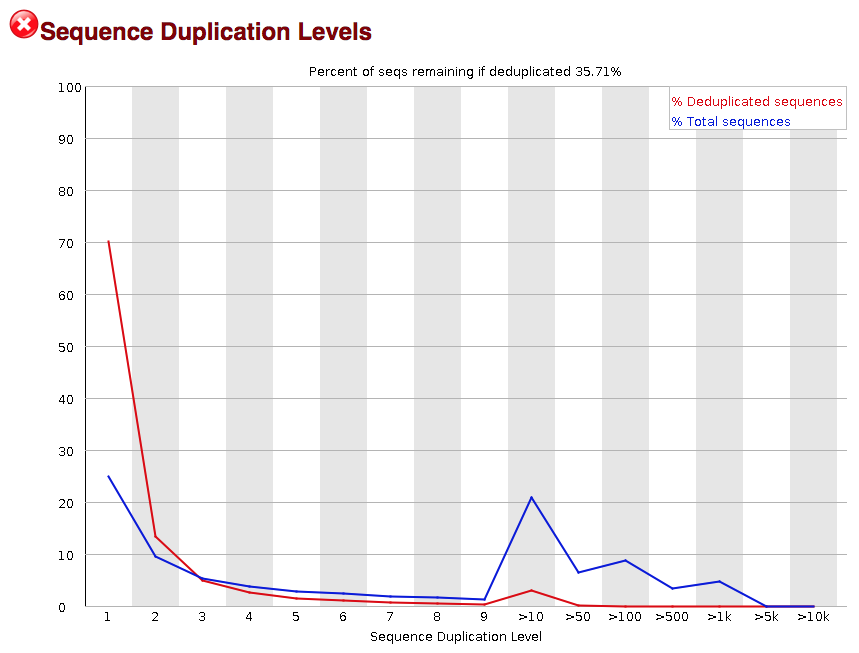
Quality control: Assessing FASTQC results | Introduction to RNA-Seq using high-performance computing - ARCHIVED