Interplay between hydrophobicity and the positive-inside rule in determining membrane-protein topology | PNAS

Interplay between hydrophobicity and the positive-inside rule in determining membrane-protein topology | PNAS
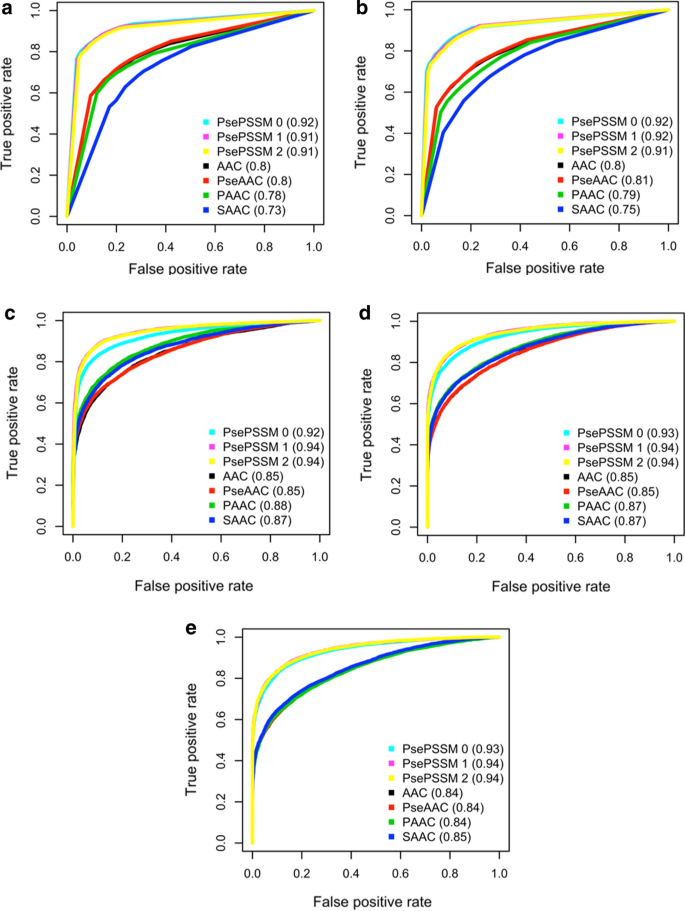
Membrane protein contact and structure prediction using co-evolution in conjunction with machine learning | PLOS ONE

Topology Prediction Improvement of α-helical Transmembrane Proteins Through Helix-tail Modeling and Multiscale Deep Learning Fusion - ScienceDirect
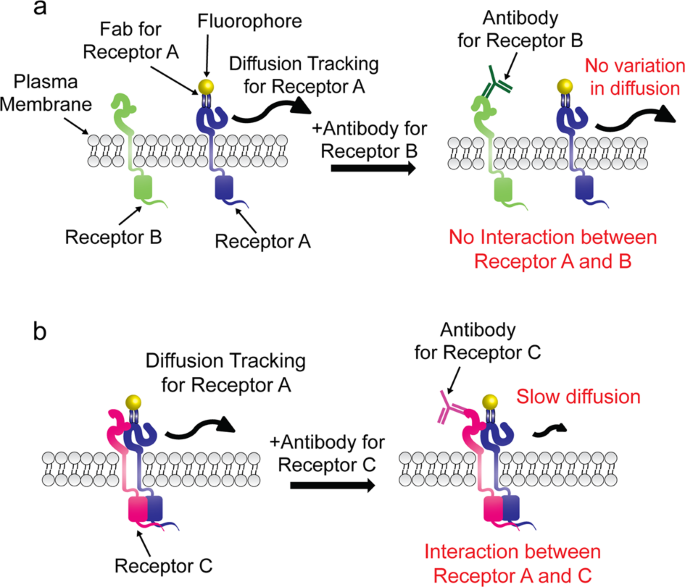
Analysis of transient membrane protein interactions by single-molecule diffusional mobility shift assay | Experimental & Molecular Medicine

MutTMPredictor: Robust and accurate cascade XGBoost classifier for prediction of mutations in transmembrane proteins - ScienceDirect

Interplay between hydrophobicity and the positive-inside rule in determining membrane-protein topology | PNAS
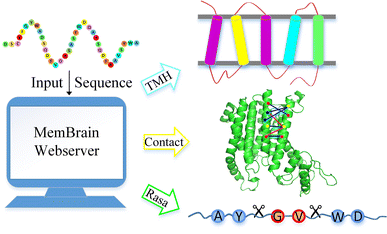
MemBrain: An Easy-to-Use Online Webserver for Transmembrane Protein Structure Prediction | SpringerLink
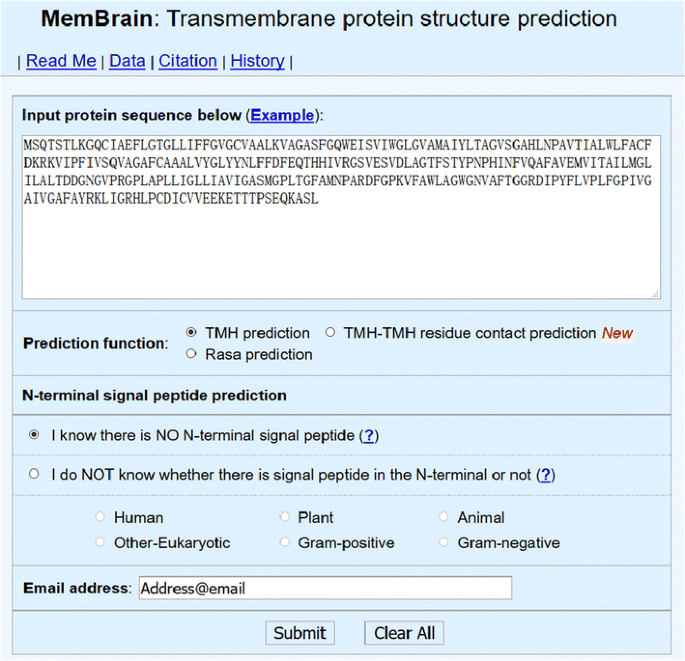
MemBrain: An Easy-to-Use Online Webserver for Transmembrane Protein Structure Prediction | SpringerLink
Membrane protein contact and structure prediction using co-evolution in conjunction with machine learning | PLOS ONE

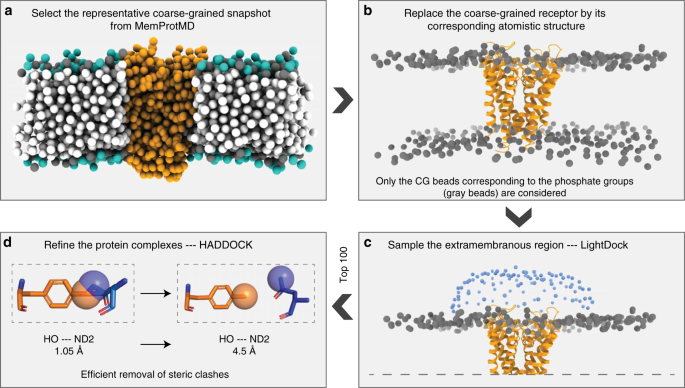
Getting to know each other: PPIMem, a novel approach for predicting transmembrane protein-protein complexes - ScienceDirect
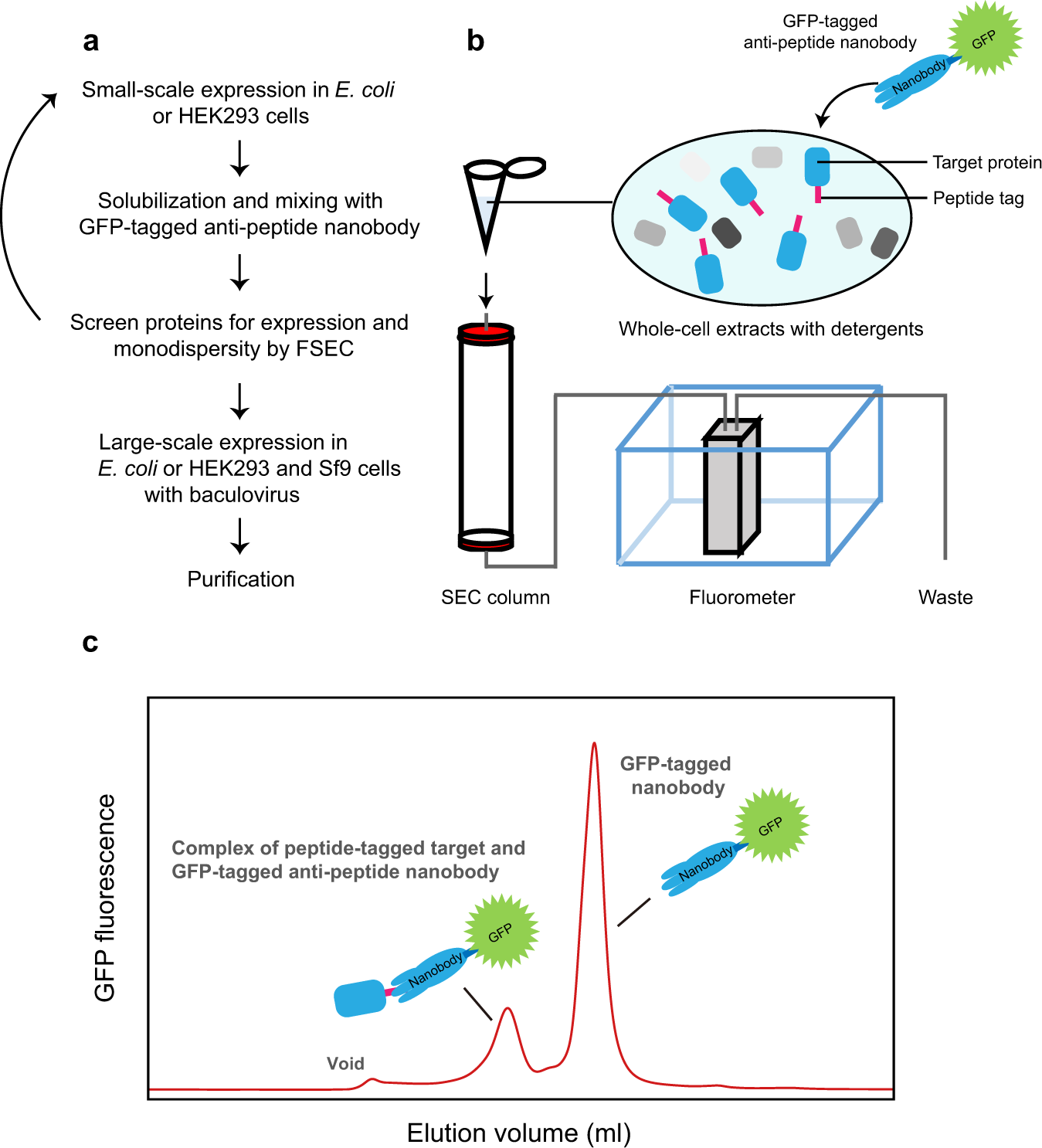
Fluorescence-detection size-exclusion chromatography utilizing nanobody technology for expression screening of membrane proteins | Communications Biology

Interplay between hydrophobicity and the positive-inside rule in determining membrane-protein topology | PNAS
A lipophilicity-based energy function for membrane-protein modelling and design | PLOS Computational Biology